Scaling the NAO's Stealth Pathogen Early-Warning System
By Jeff Kaufman 🔸 @ 2025-04-02T13:28 (+105)
This is a linkpost to https://naobservatory.org/blog/scaling-our-early-warning-system/
Summary: The NAO will increase our sequencing significantly over the next few months, funded by a $3M grant from Open Philanthropy. This will allow us to scale our pilot early-warning system to where we could flag many engineered pathogens early enough to mitigate their worst impacts, and also generate large amounts of data to develop, tune, and evaluate our detection systems.
One of the biological threats the NAO is most concerned with is a 'stealth' pathogen, such as a virus with the profile of a faster-spreading HIV. This could cause a devastating pandemic, and early detection would be critical to mitigate the worst impacts. If such a pathogen were to spread, however, we wouldn't be able to monitor it with traditional approaches because we wouldn't know what to look for. Instead, we have invested in metagenomic sequencing for pathogen-agnostic detection. This doesn't require deciding what sequences to look for up front: you sequence the nucleic acids (RNA and DNA) and analyze them computationally for signs of novel pathogens.
We've primarily focused on wastewater because it has such broad population coverage: a city in a cup of sewage. On the other hand, wastewater is difficult because the fraction of nucleic acids that come from any given virus is very low,[1] and so you need quite deep sequencing to find something. Fortunately, sequencing has continued to come down in price, to under $1k per billion read pairs. This is an impressive reduction, 1/8 of what we estimated two years ago when we first attempted to model the cost-effectiveness of detection, and it makes methods that rely on very deep sequencing practical.
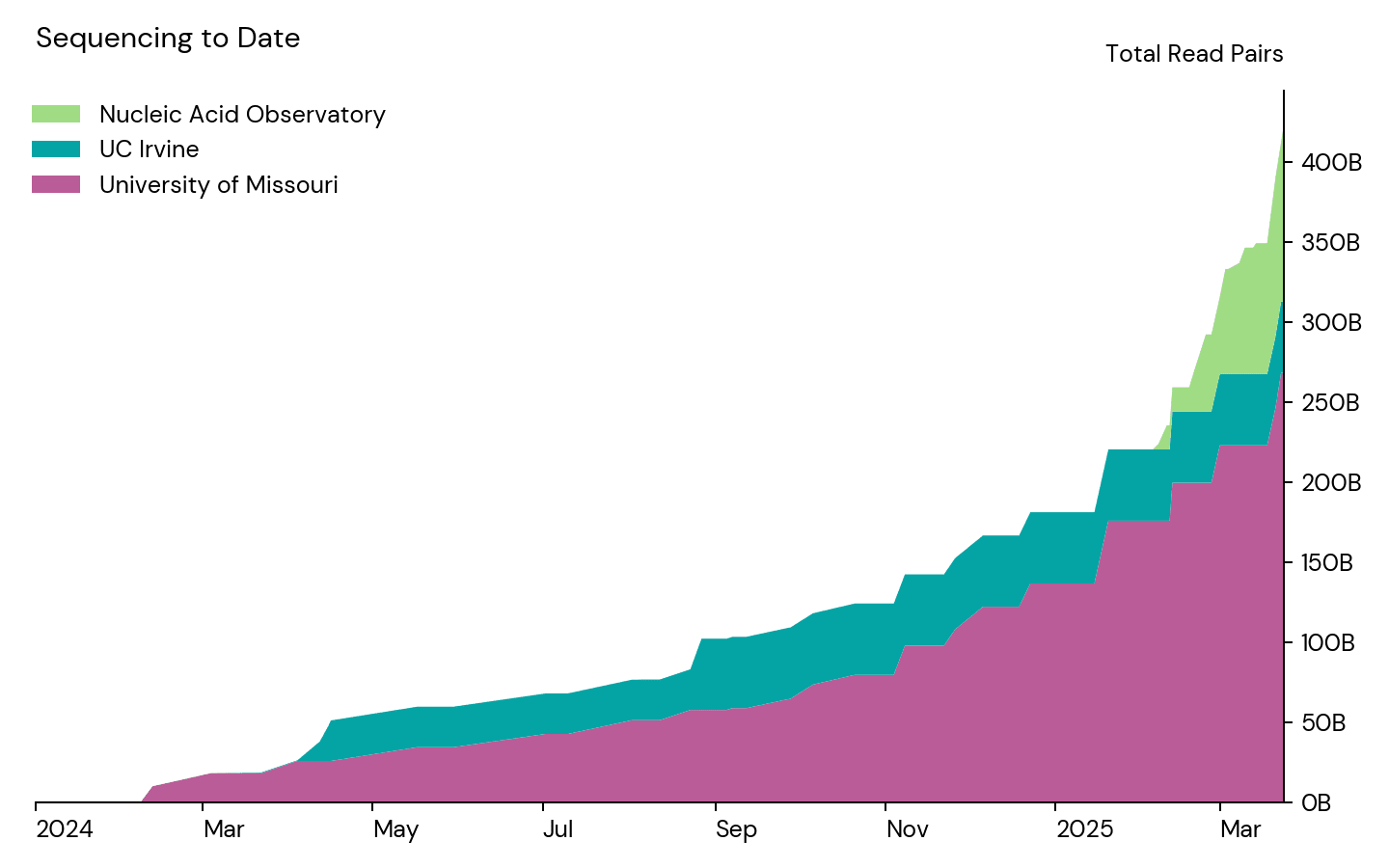
Over the past year, in collaboration with our partners at the University of Missouri (MU) and the University of California, Irvine (UCI), we started to sequence in earnest:
We believe this represents the majority of metagenomic wastewater sequencing produced in the world to date, and it's an incredibly rich dataset. It has allowed us to develop and test our algorithms for pathogen identification, and we're eager to share it with others who are working to develop their own computational approaches to this problem. This is a valuable start, and is enough to provide very sensitive coverage of gastrointestinal viruses. To get a pilot early warning system to where it could usefully flag other viruses, however, we'll need to ramp up sequencing substantially.[2]
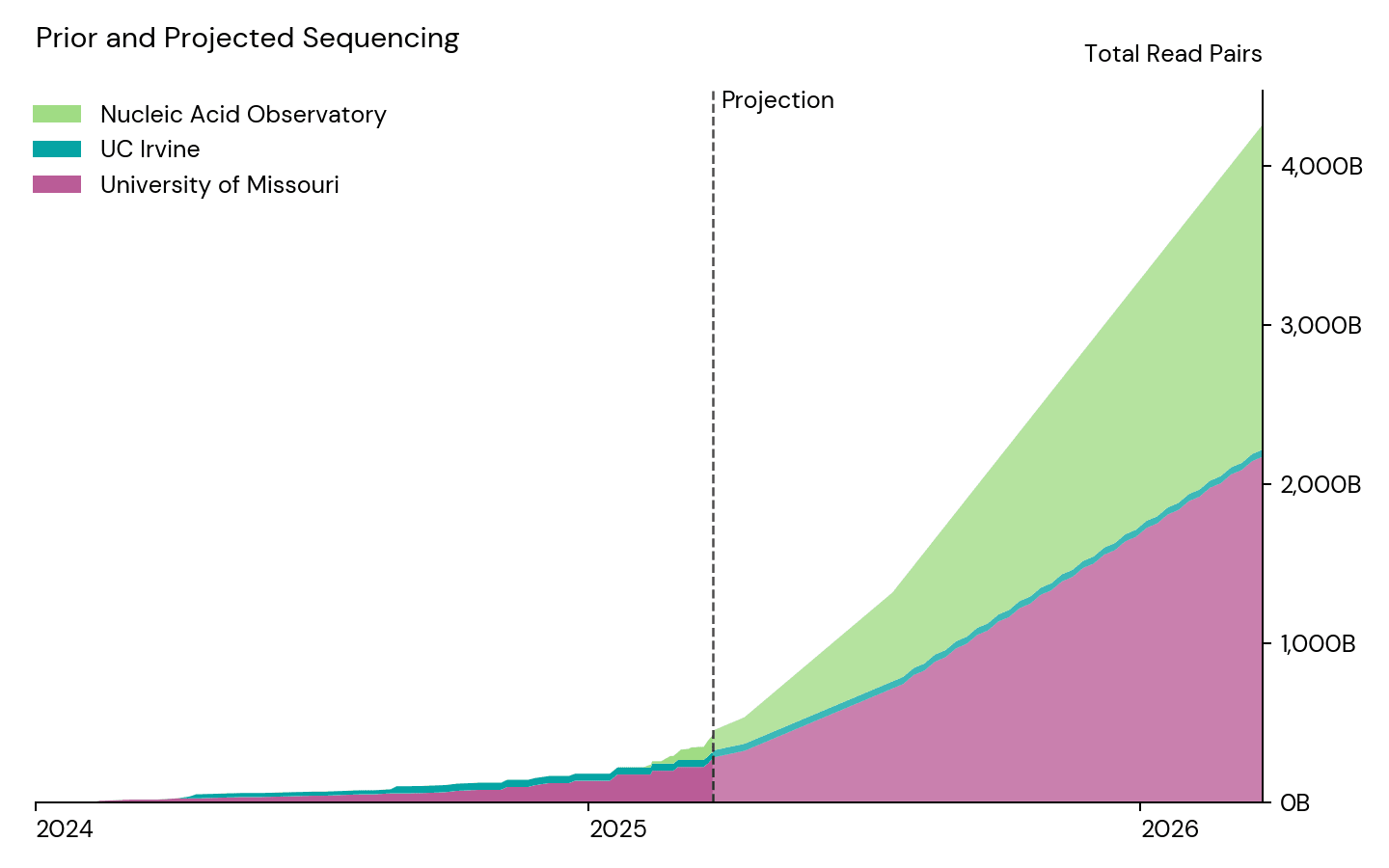
To this end, we're pleased to share that Open Philanthropy has granted $3M to the NAO over one year to fund a significant scale-up of our wastewater sequencing, targeting three NovaSeq X 25B runs weekly. We're planning to deploy these funds both in-house and at MU:
We expect this data to be valuable for a wide range of purposes, including:
-
Using our existing methods to look for engineered viruses that might be spreading now. While we think such a virus is unlikely to be at large now or over the next year, the consequences of one spreading undetected could be very serious.
-
Developing additional methods, both reference-based and reference-free, to identify novel pathogens.
-
Providing a more general supplement to traditional PCR-based public health wastewater surveillance, dramatically expanding the number of pathogens that can be tracked.
-
Allowing researchers studying the sewer microbiome to better characterize its enormous complexity.
We are grateful for Open Philanthropy's support, and also to the hard work by our collaborators at MU. Additionally, this project wouldn't be possible without our partners around the country, in academic labs and treatment plants, who are providing wastewater samples. We're very excited about what this increased scale will allow us to accomplish over the next year, and as always, we're excited to collaborate with others who are thinking along similar lines.
Even with centrifugation and filtration to enrich for viral particles, you still have to look through a lot of gut and sewer bacteria to find any. While it's possible to use targeted methods, such as hybrid capture probes that pull out sequences that match specific known pathogens, a system based on this approach would fail to detect pathogens that didn't match the probes. Instead, the NAO has worked to optimize sequence-agnostic enrichment methods, and to handle the poor signal-to-noise ratio with sufficiently deep sequencing. ↩︎
For example, we estimate that to flag a pathogen that sheds like Influenza A before about 2% of people in the monitored sewersheds have been infected, in the typical case, you'd need three NovaSeq X 25B runs each week. ↩︎
NickLaing @ 2025-04-02T14:36 (+10)
Perhaps silly question as you've probably written about this before, have you tried getting people to (with you blinded) dump both natural and engineered DNA in wastewater in different quantities at random times to see how good your system is at picking it up?
Jeff Kaufman 🔸 @ 2025-04-02T15:51 (+15)
Not a silly question, and not something where I think we've talked about plans publicly yet. Some sort of red-teaming is something I'd like to see us do in the second half of 2025. Most likely starting with fully computational spike-ins (much cheaper, faster to iterate on) and then real engineered viral particles.
Angelina Li @ 2025-04-02T16:57 (+6)
Wow, this is so exciting!! Thanks for sharing, and congratulations team!
To this end, we're pleased to share that Open Philanthropy has granted $3M to the NAO over one year to fund a significant scale-up of our wastewater sequencing, targeting three NovaSeq X 25B runs weekly.
Wow 😍. That's great. And if I read footnote 2 right, the implication is that by end of 2025, you'd aim to be able to detect a pathogen that sheds like Influenza A in cities you monitor before 2% of the population is infected? Or is that not quite right because you're targeting 3 such runs weekly across all cities (maybe I should say "sewersheds"?) so you wouldn't quite be able to hit that point yet?
I had some other basic / not-an-expert questions but no pressure to engage :)
- Which cities are you monitoring again?
- It sounds like from this notebook you're still trying to figure out how valuable monitoring one city is from the perspective of catching any global pandemic, so I assume one weakness of this approach is in the geographic restrictions. Although I've vaguely heard of wastewater monitoring in a network of airports / aircrafts as a way to get around this (I can't tell if that's just an idea right now or if it's already being implemented, though.)
- Was the 2% threshold chosen for a particular reason?
Jeff Kaufman 🔸 @ 2025-04-02T17:53 (+8)
if I read footnote 2 right, the implication is that by end of 2025, you'd aim to be able to detect a pathogen that sheds like Influenza A in cities you monitor before 2% of the population is infected?
Yes, that's right. Though sensitivity in practice could be higher or lower:
-
As we gather more data we'll get a better understanding of how easy or hard it is to detect Influenza A, along with other pathogens. Our influenza estimates are based on ~300 observations, but we now have the data to estimate for the 2024-2025 flu season with a lot more data. This is mostly a matter of someone taking the time to dig into it and put out an updated estimate.
-
We're still trying to increase sensitivity:
- Testing better wet lab methods
- Getting pooled airplane lavatory samples again, which have a ~20x higher human contribution
- Figuring out which municipal sewersheds have the highest human contribution and focusing there
-
The projection is based on an assumption of 9d end to end time, and is relatively sensitive to timing: if your pathogen doubles every 3d then the difference between a 9d and 12d turnaround time is 2x sensitivity. We're currently well above 9d, but we're on a track to get to ~7d via agreements with sequencing machine operators to reserve capacity and streamlining our processes. And then there are more expensive ways to get down to ~4d with serious investment in logistics (buy your own sequencer, run it daily, use the 10B flow cell for faster turnarounds, lab runs around the clock).
Which cities are you monitoring again?
Chicago IL, Riverside CA, and several others we hope to be able to name publicly soon.
I assume one weakness of this approach is in the geographic restrictions. Although I've vaguely heard of wastewater monitoring in a network of airports / aircrafts as a way to get around this (I can't tell if that's just an idea right now or if it's already being implemented, though.)
Yes, that's a real issue. Cosmopolitan US cities are not terrible from this perspective, especially if you have a bunch with different international connections, but they're still not good enough. Airplane lavatory sampling would be much better, not just because of this issue but also because (as I mentioned briefly above) they're much higher quality samples. We're working on this, but it's much more difficult than bringing on municipal treatment plant partners.
Was the 2% threshold chosen for a particular reason?
No, it's that 3x 25B is about the most we're able to scale to at this stage. If we thought we could manage the scale 1% would have probably been our target, though 1% is still pretty arbitrary. Lower is better, since that means mitigations are more effective when deployed, but cost goes up dramatically as you lower your target.
NickLaing @ 2025-04-02T18:40 (+6)
Do you think 1% is very useful in practise? That seems very high to me and I would have thought by that stage we would know through other means already? Or is the plan to lower the threshold as the tech improves and aim for something lower?
Jeff Kaufman 🔸 @ 2025-04-02T20:11 (+6)
I agree 1% high, and I wish it were lower. On the other hand, we're specifically targeting stealth pathogens: ones where any distinctive symptoms come well after someone becomes contagious. Absent a monitoring system, you could be in a situation most people had been infected before anyone noticed there was something spreading. Flagging this sort of pathogen at 1% cumulative incidence still gives some time for rapid mitigations, though it's definitely too late to nip it in the bud.
Angelina Li @ 2025-04-02T17:44 (+6)
From this post:
They’re now sequencing wastewater from eight sewersheds across four metropolitan areas, with the addition of Riverside CA (in collaboration with Jason Rothman) in December.
In Fall 2023 we partnered with CDC’s Traveler-based Genomic Surveillance program and Ginkgo Biosecurity to collect and sequence both pooled airplane lavatory waste and municipal wastewater influent and sludge. We’ve submitted a full set of aliquots to MIT’s BioMicroCenter for high-throughput library preparation, and will be sending the libraries to Broad Clinical Labs for sequencing later this quarter.
I see, that answered some of my questions. I still feel confused how big a sewershed is relative to a city, and how much that matters from the perspective of early detection. But no pressure to engage, was just curious. Exciting!
Jeff Kaufman 🔸 @ 2025-04-02T17:58 (+8)
I still feel confused how big a sewershed is relative to a city
A sewershed can vary dramatically in size: it's the area that drains to some collection point (generally a treatment plant) and different cities are laid out differently. I'm most familiar with Boston (after refreshing the MWRA Biobot Tracker intently during COVID-19) and here the main plant serves ~2M people divided between the North and South systems:
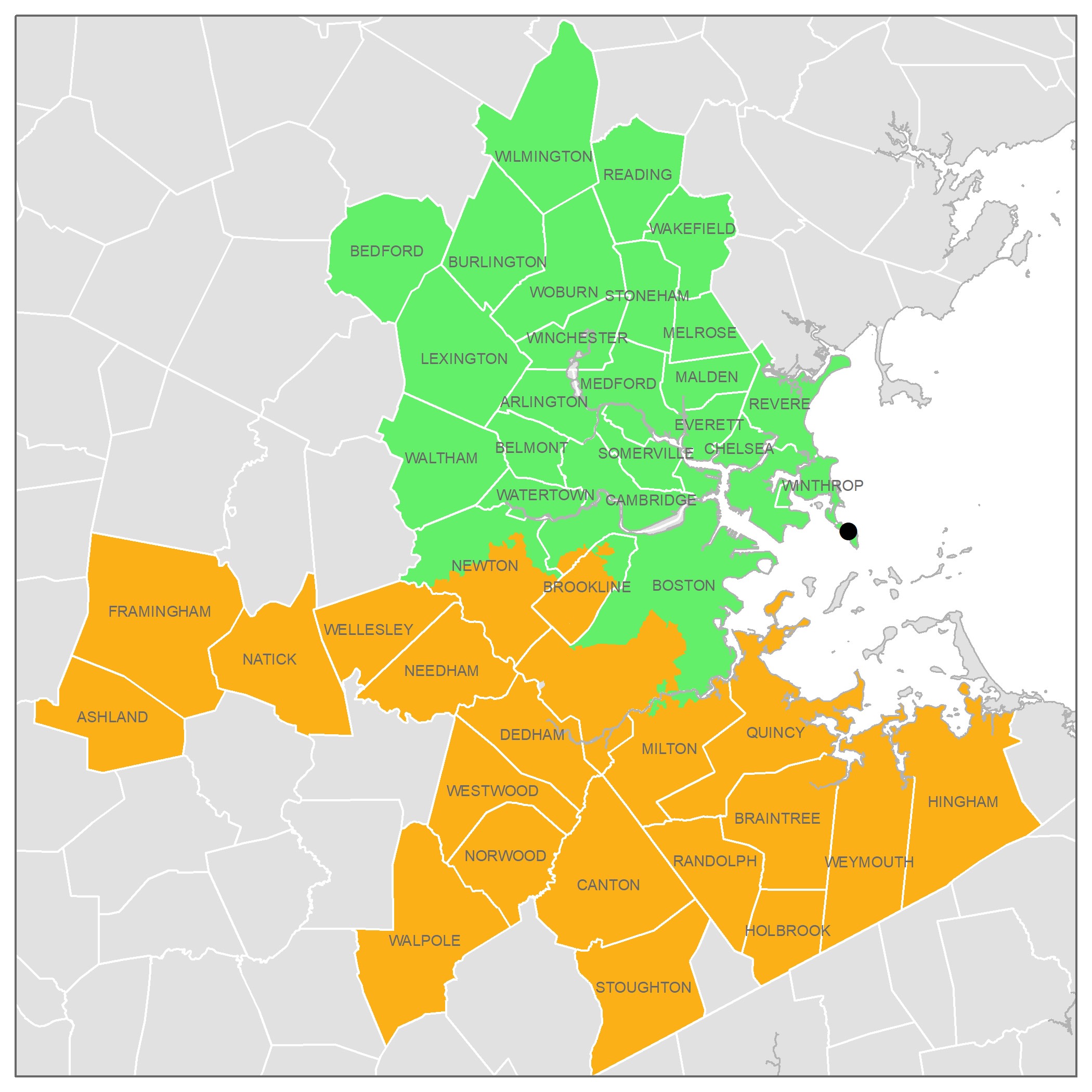
Some other cities have much smaller plants (and so smaller sewersheds), a few have larger ones.
We're not sure yet about the effect of size. It's possible that small ones are better because the waste is 'fresher' and you spend fewer of your observations (sequencing reads) on bacteria that replicates in the sewer. Or it's possible that larger ones are better because they can support more observations (deeper sequencing).
Angelina Li @ 2025-04-02T18:02 (+6)
Nice. You're such a fast writer! Very helpful, thank you!
Jeff Kaufman 🔸 @ 2025-04-02T18:18 (+6)
It helps that I'm writing about stuff we've discussed internally a lot! Thanks for the good questions!